deepDTnet receives provisional patent to accelerate drug repurposing and minimize the translational gap in development
Although conventional drug development aims to design drugs that selectively target a single molecular entity (e.g., a disease-driving protein), drugs often are found to interact with more than one target. These off-target interactions can be problematic as they may result in adverse effects and suboptimal drug effectiveness. However, therapeutics with multiple targets provide opportunities for repurposing drugs to treat diseases that do not yet have effective therapies, as long as the molecular targets with which a drug will interact can be comprehensively identified.
Advertisement
Cleveland Clinic is a non-profit academic medical center. Advertising on our site helps support our mission. We do not endorse non-Cleveland Clinic products or services. Policy
Novel drug-target interactions (DTIs) can be determined via experimentation, but it remains a costly and time-consuming process. Feixiong Cheng, PhD, Genomic Medicine Institute, is working to develop a more streamlined and cost-effective approach that integrates artificial intelligence (AI) and network medicine technologies.
In a recent cover paper published in Chemical Science, a team of researchers led by Dr. Cheng developed a computational methodology that utilizes heterogeneous data on known DTIs to predict new interactions with greater accuracy than previous methods. Known as deepDTnet, this methodology embeds a network connecting drugs, targets and diseases and, via deep learning (i.e., a type of machine learning in AI), infers the targets with which a drug will interact.
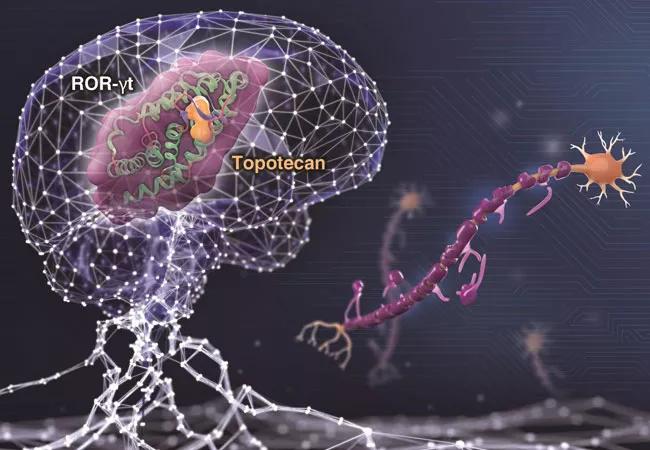
To validate deepDTnet for drug repurposing, Dr. Cheng’s team used deepDTnet to predict drugs that would interact with ROR-γt (retinoic acid receptor-related orphan receptor-gamma t), a protein involved in pro-inflammatory processes associated with autoimmune diseases, such as multiple sclerosis. Experimental testing found that, among the 18 deepDTnet-predicted drugs, six had human ROR-γt inhibitory activities greater than 30%, with the approved chemotherapy drug topotecan demonstrating a direct ROR-γt inhibitor (IC50=0.43 µM) and a potential therapeutic effect in a preclinical model (experimental autoimmune encephalomyelitis) of multiple sclerosis.
These findings indicate that deepDTnet offers a powerful network-based deep learning methodology for target identification to accelerate drug repurposing and minimize the translational gap in drug development. They have applied a provisional U.S. patent for this new technology. Furthermore, they demonstrate the value of methods that integrate AI and network medicine technologies to systematically identify novel targets and repurposable drugs for multiple human complex diseases – a concept that Dr. Cheng’s team continues to explore. For instance, in a study published in Bioinformatics, they present an improved network-based computational framework called AOPEDF (arbitrary-order proximity embedded deep forest) that builds upon deepDTnet and offers an effective tool for accelerating target-centered drug repurposing and therapeutic development for understudied diseases.
Advertisement
Figure: A network-based deep learning methodology, termed deepDTnet, for novel target identification and in silico drug repurposing. This image was prepared by Cleveland Clinic Center for Medical Art & Photography.
Advertisement
Advertisement

First full characterization of kidney microbiome unlocks potential to prevent kidney stones

Researchers identify potential path to retaining chemo sensitivity
Large-scale joint study links elevated TMAO blood levels and chronic kidney disease risk over time
Investigators are developing a deep learning model to predict health outcomes in ICUs.

Preclinical work promises large-scale data with minimal bias to inform development of clinical tests

Cleveland Clinic researchers pursue answers on basic science and clinical fronts

Study suggests sex-specific pathways show potential for sex-specific therapeutic approaches

Cleveland Clinic launches Quantum Innovation Catalyzer Program to help start-up companies access advanced research technology