Using data science to identify intrinsic patterns in cancer data
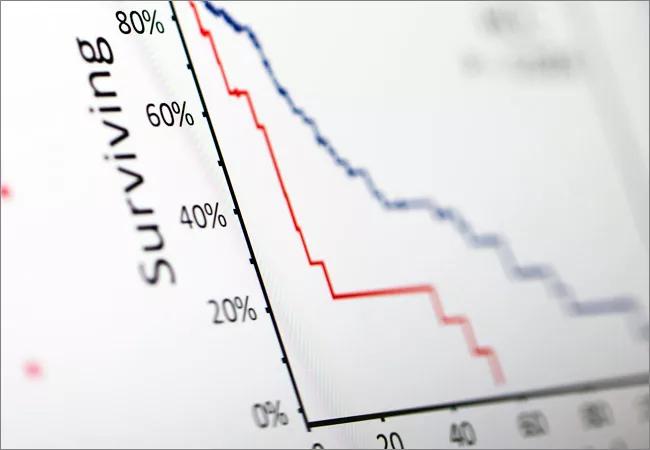
Quantitative analysis and pattern recognition using mathematical models is necessary to bring clarity to complex data and, thereby, pivotal to precision medicine in oncology.
Advertisement
Cleveland Clinic is a non-profit academic medical center. Advertising on our site helps support our mission. We do not endorse non-Cleveland Clinic products or services. Policy
Brian Hobbs, PhD, an Associate Staff in Cleveland Clinic’s Department of Quantitative Health Sciences and Section Head of Cancer Biostatistics, weighed in on the utility of quantitative analysis and mathematical modeling in cancer research, as well as future prospects in this field.
How has the definition of precision medicine in oncology changed over the years and how has this change affected oncology data analysis?
Oncology therapies have evolved over time, from cocktails of cytotoxic drugs with the potential for radiotherapy and surgery, to non-cytotoxic therapies that target specific pathways in tumor cells or promote anti-cancer immunity.
Precision medicine, in essence, aims to understand treatment benefit heterogeneity at the patient/tumor level and establish actionable targets through biomarker-guided therapies. Specific targets and treatment strategies themselves are as varied as the biologies of cancer subtypes. As an objective, precision medicine has perhaps not deviated from its original state. What has evolved, however, is the complexity with which patient subpopulations are defined and interrogated. Ten years ago, precision medicine mostly focused on classifying patients into biomarker positive or biomarker negative cohorts.
Advances in biology and immunology continue to refine our understanding of cancer. The idea that tumor biology and/or host immunity may better define populations than cancer histology and/or clinical site has evolved and defined new research programs and study designs such as basket, umbrella, and platform clinical trial designs. We have to really think about the subtypes of patients based on many features that could be present or co-exist together in the tumor microenvironment, including genetic alterations of the cells, existing clinical comorbidities, prior treatment history, effects of combination therapies, etc. With this, the role of data science becomes central to the endeavor. Specifically, we contribute through application of the inferential process to either validate or discover potential mechanisms of tumor cell growth, survival, angiogenesis, and systematic suppression of cancer immunity.
Advertisement
Where does your work fit in the overall goal of personalized cancer treatment delivery?
Our work is interdisciplinary in nature — it involves collaboration with clinical investigators, computer (data) scientists, and translational scientists. Our job is to facilitate a deeper understanding of data through dimension reduction with mathematical models that identify patterns; these models are devised to characterize the patterns as well as describe the extent of evidence for their existence beyond chance in a manner that can be disseminated objectively to the research community and tested with future study.
We work by structuring complex data and devising mathematical models that reflect the relationships among data. An example is a high-profile paper published recently in Scientific Reports and based on our work done at MD Anderson. In this study, which focused on a subset of patients treated with definitive surgery for NSCLC, we were able to subtype a patient’s immune response (the innate reaction to the presence of a malignant mass) based on the radiographic textural patterns we identified from the pre-treatment CT images. Moreover, we defined the extent of prognostic heterogeneity that may be attributable to the subtypes which defines the expected survival of patients presenting various patterns. This has the potential to extend the subtyping of immune phenotypes to non-invasive biomarkers.
Although most people think that our work begins at the end of a study, at the stage of data analysis, we also play a fundamental role in planning the design of a future study. We help clinical researchers plan what type of data to acquire and how to acquire it, so that it may be better understood upon study completion.
Advertisement
How do you approach the decision about which mathematical model to apply to a certain data set?
While every research project presents unique attributes and challenges, the foundation for a given analytical model, and manner in which dimension reduction can be used to identify patterns, is defined by the object of interest for inference and the framework by which informatics can be observed. Spatially orientated textural pattern data is very different from observing a sequence of durations that define the extent of time during which a patient maintained a durable response to a particular treatment regime.
We think at a general “structure-level” to engineer analytical tools and concepts to solve specific problems.
In which directions do you see the field of biostatistics developing in the future?
There is no clear boundary between the fields of biostatistics and bioinformatics. I think that we will see a continued shift in the emphasis from simply acquiring data to understanding the patterns among data in both fields. As a result, the scope of our work will magnify tremendously as we gain access to more structured informatics that necessitate new models for signal identification. It’s my expectation that the demand for quantitative researchers in biomedicine, oncology in particular, will continue to expand with the emergence and widespread adoption of precision medicine.
Advertisement
Advertisement

First full characterization of kidney microbiome unlocks potential to prevent kidney stones

Researchers identify potential path to retaining chemo sensitivity
Large-scale joint study links elevated TMAO blood levels and chronic kidney disease risk over time
Investigators are developing a deep learning model to predict health outcomes in ICUs.

Preclinical work promises large-scale data with minimal bias to inform development of clinical tests

Cleveland Clinic researchers pursue answers on basic science and clinical fronts

Study suggests sex-specific pathways show potential for sex-specific therapeutic approaches

Cleveland Clinic launches Quantum Innovation Catalyzer Program to help start-up companies access advanced research technology