Better assessing individual risk of recurrence
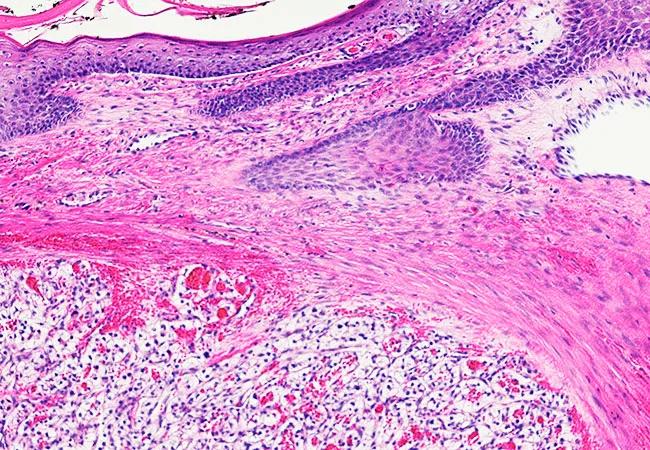
Among patients with clinically localized renal cell carcinoma (RCC), 30% will relapse with distant metastatic disease after nephrectomy. The risk of recurrence influences the intensity of follow-up imaging, the guidance provided to patients, and consideration of adjuvant treatment and clinical trials.
Advertisement
Cleveland Clinic is a non-profit academic medical center. Advertising on our site helps support our mission. We do not endorse non-Cleveland Clinic products or services. Policy
Predicting recurrence can be challenging: The commonly used prognostic models, such as SSIGN and UISS, consider only clinicopathologic variables such as T stage and grade and do not capture tumor biology or adequately discriminate recurrence risk. They are also subject to varied interpretations by observers.
Over the past decade, multigene assays have been developed for RCC that provide information beyond traditional parameters in multiple tumor types that can better assess individual risk. At the 2019 Genitourinary Cancers Symposium, Brian Rini, MD, leader of Cleveland Clinic’s Genitourinary Program, provided an overview of genetic prognostic models for RCC.
A multigene signature based on cell cycle proliferation (CCP) was developed based on 565 patients at Massachusetts General Hospital and the University of Michigan who underwent radical nephrectomy for localized phase T1-T3 clear cell, papillary or chromophobe RCC.
RNA was extracted from the highest grade area of the viable tumor, and CCP score was calculated based on the unweighted average of 31 cell cycle genes normalized to the expression of 15 housekeeping genes. The CCP score was compared with the Karakiewicz nomogram, and a composite (R-CCP) score was developed.
The CCP score was an independent predictor of recurrence (hazard ratio [HR] 1.50, 95% confidence interval [CI], 1.07-2.09) and disease-specific mortality (HR 2.49, 95% CI, 1.53-4.04). Combining the molecular classifier with baseline clinical variables allowed for accurate, patient-specific risk assessment to guide clinical management.
Advertisement
ClearCode34, a 34-gene classifier for assigning tumors to good risk (ccA) and poor risk (ccB) subtypes, was based on 72 clear cell RCC sample standards. The classifier was applied to RNA-sequencing data from 380 nonmetastatic clear cell RCC samples from The Cancer Genome Atlas and 157 clinical samples collected at the University of North Carolina.
The ccA tumors overexpressed genes associated with hypoxia, angiogenesis, fatty acid metabolism and organic acid metabolism. The ccB tumors overexpressed genes that regulate epithelial to mesenchymal transition, the cell cycle and wound healing.
ClearCode34 had a hazard ratio for disease-specific mortality of 2.9. This model provides prognostic stratification that improved upon established algorithms to assess risk for recurrence and death for nonmetastatic clear cell RCC patients.
In developing this assay, 732 genes from published data and biologic pathways that are functionally important in clear cell RCC were analyzed for their association with clinical outcomes in 942 Cleveland Clinic patients with stage I-III clear cell RCC who had undergone a nephrectomy. A total of 516 genes were associated with recurrence-free interval, and 11 were selected by further statistical analyses and combined with five reference genes to develop a recurrence score algorithm.
The 16-gene recurrence score was initially validated in an independent cohort of 626 patients with stage I-III clear cell RCC who had also undergone nephrectomy. In multivariate analyses, the recurrence score was significantly associated with the risk of tumor recurrence (HR per 25-unit increase in the score 3.37 [95% CI, 2.23-5.08]). It was able to identify a clinically significant number of both high-risk stage I and low-risk stage II-III patients.
Advertisement
In a second validation study, the 16-gene assay was evaluated in 212 patients, with a focus on 193 with stage III RCC, from the S-TRAC randomized trial of adjuvant sunitinib. The 16-gene recurrence score results predicted time to recurrence, disease-free survival and renal cancer-specific survival in both arms, with the strongest results observed in the placebo arm. When high versus low recurrence score groups were compared, the HR for recurrence was 9.18 (95% CI, 2.15-39.24) in the placebo arm. The study provided 1B evidence for the 16-gene assay, offering the strongest evidence base of the three genetic models.
Genetic models have the potential to refine the risk of recurrence and guide physician management and physician and patient decisions concerning adjuvant treatment. As yet, they have not been used in clinical settings. Based on the evidence to date, Dr. Rini recommends further investigation of genetic models for predictive value, utility in adjuvant IO trials and other settings, such as SRM.
Advertisement
Advertisement

Radiation therapy helped shrink hand nodules and improve functionality

Standard of care is linked to better outcomes, but disease recurrence and other risk factors often drive alternative approaches

Phase 1 study demonstrates immune response in three quarters of patients with triple-negative breast cancer

Multidisciplinary teams bring pathological and clinical expertise

Genetic variants exist irrespective of family history or other contributing factors

Study shows significantly reduced risk of mortality and disease complications in patients receiving GLP-1 agonists

Structured interventions enhance sleep, safety and caregiver resiliency in high-acuity units

Addressing rare disease and challenging treatment course in an active young patient