Toward detailed, personalized maps of identified and interpreted genomic variation
Thanks to remarkable scientific and technological advancements, researchers now have a deluge of sequencing data, which, with the right analysis, may help with the discovery and development of targeted cancer treatments. In a cover article published in Pharmacological Reviews, Cleveland Clinic’s Feixiong Cheng, PhD, Genomic Medicine, and collaborators review the current use of personal mutanomes in the discovery of modern oncology drugs, including therapies that are targeted to specific genomic or molecular profiles, as well as immunotherapies.
Advertisement
Cleveland Clinic is a non-profit academic medical center. Advertising on our site helps support our mission. We do not endorse non-Cleveland Clinic products or services. Policy
Although we know that cancers are complex with multiple alterations that differ greatly from person to person, even the current therapies aim at a single target. Dr. Cheng and colleagues propose a systems-wide approach to look for the sum total of all relevant alterations, so called systems medicine, on a person-to-person basis.
According to Dr. Cheng, “at the root of precision oncology is the hypothesis that cancer treatment would be considerably better if therapies were guided by a tumor’s genomic alterations.” Such molecularly targeted agents suppress only the pathways that fuel cancer cells by inhibiting the function of orthosteric and allosteric sites.
Dr. Cheng describes personal mutanomes as detailed maps of identified and interpreted genomic variation. In this case, three-dimensional models illuminate the impact of treatments developed specifically as a result of mutations discovered when the tumor was diagnosed. With the help of computational biophysics, mutanomes can identify precisely which proteins or protein networks to target with drug therapies, with fewer or no side effects on the surrounding structures. Dr. Cheng proposes “a personal mutanome, an emerging personalized medicine infrastructure, for accelerating the development of precision oncology.”
The personal cancer mutanome infrastructure has four core components: performing tumor genetic and genomic testing; using bioinformatics or computational biology tools to identify actionable biomarkers; preclinical validation; and guiding mono- or combination therapies. The group applied this process in a breast cancer case study to illustrate the exciting potential of these developments.
Advertisement
Moving from the personal mutanome to precision medicine poses challenges for all healthcare stakeholders, according to Dr. Cheng, including:
Dr. Cheng urges academic institutions and pharmaceutical industries to work together to bring more affordable, personalized medications to patients with cancer and other diseases. The personal mutanome is an example of an innovative computational tool that models the DNA landscape of a patient’s genome, which may help clinicians to fulfill the promise of precision medicine by providing the right drugs, at the right dose, to the right patient at the right time.
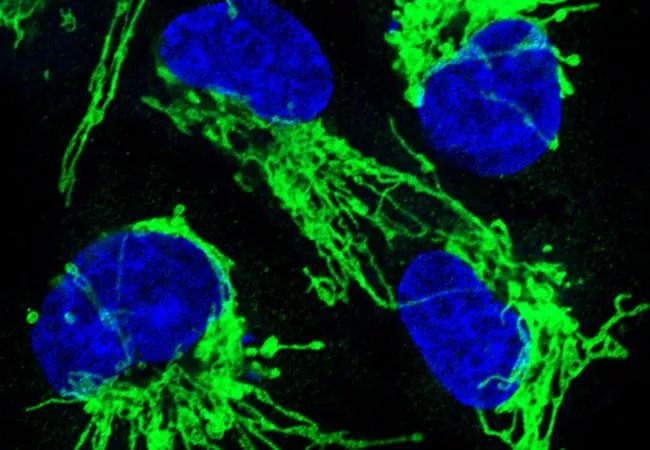
Image: Ras-driven cancer. About a third of all human cancers, including a high percentage of pancreatic, lung, and colorectal cancers, are driven by mutations in RAS genes. When Ras genes are mutated, cells grow uncontrollably and evade death signals. Ras mutations also make cells resistant to some therapies. Many of these effects occur as a result of Ras-induced changes in mitochondrial shape. This image shows mitochondrial staining (green) and nuclear staining (blue) following inhibition of the mitochondrial fission machinery in human epithelial cells engineered to express oncogenic Ras. Source: NCI Visuals Online.
Advertisement
Advertisement

First full characterization of kidney microbiome unlocks potential to prevent kidney stones

Researchers identify potential path to retaining chemo sensitivity
Large-scale joint study links elevated TMAO blood levels and chronic kidney disease risk over time
Investigators are developing a deep learning model to predict health outcomes in ICUs.

Preclinical work promises large-scale data with minimal bias to inform development of clinical tests

Cleveland Clinic researchers pursue answers on basic science and clinical fronts

Study suggests sex-specific pathways show potential for sex-specific therapeutic approaches

Cleveland Clinic launches Quantum Innovation Catalyzer Program to help start-up companies access advanced research technology