$14M NIH grant supports research toward medications to prevent progression

Image content: This image is available to view online.
View image online (https://assets.clevelandclinic.org/transform/4146f135-298c-4c22-b274-26ee4bee3191/Heart_Chung_129483_09-22-17_0020_TM_hrvrtcc-scaled-jpg)
Mina Chung, MD
The prevalence of atrial fibrillation (AF) in the U.S. is projected to double by 2030, largely due to aging of the population and obesity. Few effective therapies exist for AF, however, and no new pharmacologic agents have been approved for the condition in more than 10 years.
Advertisement
Cleveland Clinic is a non-profit academic medical center. Advertising on our site helps support our mission. We do not endorse non-Cleveland Clinic products or services. Policy
Building on nearly 20 years of research on AF, clinicians and scientists in Cleveland Clinic’s Heart, Vascular & Thoracic Institute and Lerner Research Institute are hoping to fill that gap through four synergistic projects that have their roots in — but go well beyond — genetics.
Funded by a five-year, $14.2 million National Institutes of Health grant, the work by the multidisciplinary team is designed to advance existing genomics — and omics more broadly — and develop a pipeline of repurposed drugs that can prevent progression of AF in adults.
To achieve that goal, the researchers will address gaps in five areas of knowledge:
“The need for novel targets for AF is substantial, but genetic findings so far have failed to yield clinically actionable results,” says project lead Mina Chung, MD, a cardiologist in Cleveland Clinic’s Section of Cardiac Electrophysiology and staff in the Department of Cardiovascular and Metabolic Sciences. “Our team’s goal is to bring basic research on genomic mechanistic discoveries in AF to the bedside by identifying mechanisms and targets underlying the condition that may lead to personalized therapies.”
Advertisement
More than 100 genetic loci relevant to AF have been identified since AF’s heritable nature and genetic basis were confirmed by the first GWA study (GWAS) in 2007. The locus with the strongest association with AF is near PITX2, which codes for the PITX2 transcription factor expressed early in development. PITX2 appears to suppress a left atrial sinus node program and is involved with formation of the pulmonary veins.
“PITX2 is a tantalizing candidate gene because of its association with the pulmonary veins, which are the target of ablation,” Dr. Chung explains. “Patients with AF may be born with a susceptibility to the condition, but it may manifest only in adulthood after exposure to stressors such as aging and obesity.”
Using human tissue samples from the pulmonary vein and left atrial appendage, Cleveland Clinic researchers are studying the leading pathways implicated by candidate AF risk genes. They have identified novel upstream pathways using transcriptomics and identified genetic effects on nearby gene expression. That has led them to propose — and pursue via the NIH-supported projects — a dual risk model (Figure) that may explain AF’s association with heterogeneous cardiovascular stressors.
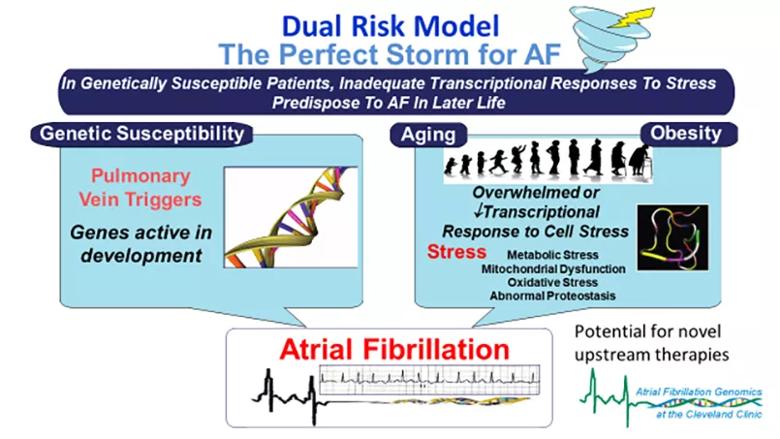
Image content: This image is available to view online.
View image online (https://assets.clevelandclinic.org/transform/3f2d099e-82c5-468a-867e-31949f4f64ee/22-HVI-3073820-CQD-Inset_jpg)
Figure. A dual risk model for atrial fibrillation (AF) proposed by Dr. Chung and colleagues in a recent review article. Reprinted, with permission, from Chung MK, et al., “Insights From Atrial Fibrillation Genomics: From Bedside to Bench and Back Again,” Cardiology in Review (2019;27[6]:302-307), https://journals.lww.com/cardiologyinreview.
Advertisement
“Thanks to our collaboration with cardiothoracic surgeons, we developed a bank of human left atrial tissues that has allowed us to make the links from DNA to mRNA to protein to function,” says Dr. Chung. “Now we will be using a network systems biology approach and layering on various omics to identify disease modules, as well as performing network-based proximity analysis to identify existing drugs that act on those modules.”
The researchers will integrate new single-nucleus RNA sequencing with AF recurrence GWAS, bulk RNA sequencing, and proteomic and metabolomic data to identify interactome-mediated gene regulatory networks underlying AF progression in the human AF tissue. The team hypothesizes that in AF, an interplay exists between systemic inflammatory, metabolic and fibrotic mechanisms and gene regulatory networks.
Understanding this interplay, they believe, is essential to gain insights into AF progression and inform therapeutic discovery. It could also be key to reducing the burden of the condition, because 25% of individuals go on to have persistent or permanent AF within five years after their initial diagnosis.
As part of the research, artificial intelligence and systems biology approaches will be applied to identify repurposable candidate drugs and drug combinations. Validation will be performed with beating cardiomyocytes derived from inducible pluripotent stem cells from healthy subjects and patients with AF. The top candidate drugs will be tested in mouse models of spontaneous AF and progression.
Advertisement
The research program’s four individual projects also aim to enhance understanding of the function of selected candidate AF risk genes; the interplay of metabolic, mitochondrial and obesity pathways in AF; and the effect of diet and the microbiome on AF. These projects ultimately will be integrated to identify drugs or drug combinations that have potential to advance to clinical trials in patients with AF.
“This is a huge team effort focused on working toward personalization of treatment and shortening of the timeline for new AF therapies,” says Dr. Chung. “This approach is necessary because the condition is vexing to many patients as they age, and AF’s impact on these individuals and our healthcare system is significant and growing.”
“We are honored to receive this significant grant from the NIH that will support our ongoing investigation of AF on multiple different levels,” adds Oussama Wazni, MD, MBA, Section Head of Cardiac Electrophysiology and Pacing. “We hope this work will expand our knowledge of the multiple pathophysiologic pathways of AF and its treatment.”
Dr. Chung’s co-investigators on the project include Jonathan Smith, PhD; David Van Wagoner, PhD; Sarah Schumacher-Bass, PhD; Sathyamangla Prasad, PhD; Robert Koeth, MD, PhD; and Julie Rennison, of the Department of Cardiovascular and Metabolic Sciences; John Barnard, PhD, of the Department of Quantitative Health Sciences; and Feixiong Cheng, PhD, of the Genomic Medicine Institute, all with Cleveland Clinic, as well as Kenneth Laurita, PhD, of MetroHealth Medical Center and Case Western Reserve University.
Advertisement
Advertisement
AVANT GUARD trial extends first-line role for ablation beyond paroxysmal atrial fibrillation
LAA closure may be compelling option in atrial fibrillation ablation patients at high risk of both stroke and bleeding
UK experts compare and contrast the latest recommendations
While results were negative for metformin, lifestyle counseling showed surprising promise
Investigational pulsed-field ablation system also yielded procedural efficiencies
Concomitant AF ablation and LAA occlusion strongly endorsed during elective heart surgery
ACC/AHA panel also upgrades catheter ablation recommendations
For the first time, risk is shown after accounting for underlying contributions of pulmonary disease